my_cache = T
knitr::opts_chunk$set(fig.pos = 'H',
cache = my_cache,
cache.path = "chache/",
dpi = 600,
fig.path = "figs/",
fig.align = "center",
fig.asp = 0.62) # goldener
library(tidyverse)
library(brms)
library(marginaleffects)
library(tidybayes)
library(ggdist)
set.seed(123)
dat <- epilepsy # get the dataset
# a simple model fitting number of epileptic seizure by the treatment (0 or 1)
mod <- brm(count ~ 1 + Trt,
family = poisson(),
data = dat,
file = "poisson_model")
theme_set(theme_bw())
# the linkfunction of the poisson regression is a log()Fun with link-functions
This site serves future me and anyone interested as a help to understand how to interpret the results of regressions that have link-functions. One of the difficulties is that the interpretation of models’ parameters changes with the link function and the type of predictors used (see this paper).
If you detect an error, please let me know: lilla.gurtner2@unibe.ch
I will illustrate this, using Bayesian regressions fitted with brms. In general, there are different packages that provide helper functions to get at the estimates of a model at the link level and at the expected and predicted response level (see this helpfull site for the distinctions). I will compare the results of three different packages: brms, tidybayes, marginaleffects.
brms has functions like posterior_linpred() and posterior_epred() that give access to the draws of the posterior estimates. They are the basis of the following packages, but the results are a bit cumbersome to work with.
tidybayes makes access to these draws simpler by providing them in a tidy format as a tibble rather than a matrix. Based on this direct access to the posterior, summary statistics can be computed / plotted.
marginaleffects uses the insight package to provide marginal means and constrasts for a given model. It provides the predictions() functions to get predictions for each observation of an existing or a synthetic data set, and the comparisons() function to take easily the difference between e.g. groups. In addition, the slopes() function gives easy access to the interpretation of the parameter estimates of continuous predictors. All three can have the prefix avg_before them to take the average of the above. A lot of nice examples for brms models can be found here.
If you are interested in the emmeans package, see this comparison with marginaleffects.
In the first section, I collect the different functions from the different packages and compare them to each other, to be best of my understanding. In the second section, I go through two examples.
Setup
An example generalized linear regression model. For the moment, we work with a single predictor with only two levels.
This is a generalized model, that can be noted as follows:
\(count \sim Poisson(lambda)\)
\(log(lambda) = Intercept + Trt * x\)
where x is a binary variable. Note that the linear part is predicting the log-transformed lambda, i.e. the log-transformed central tendency or expected value of the seizures. The linear part is thus formulated at the link-level, and to get to an expected value, it has to be back-transformed trought the inverse link-function, in this case exp().
Functions to get to the estimates
Draws on the link-level
The following functions can be used to get to the posterior samples on the link-level, that is, on the level of the linear regression term. This is, what is also returned in summary(model).
using brms package
a <- brms::posterior_linpred(mod)
glimpse(a) num [1:4000, 1:236] 2.12 2.13 2.17 2.17 2.19 ...Produces a matrix with n_samples (rows) x n_observations (cols), contains 4000 predictions on the level of the link function for each observation.
using tidybayes functions
a <- tidybayes::tidy_draws(mod)
# tibble with length = n_chains*n_draws (postwarump)
# contains the samples from the posterior in column b_Trt1 + other indicators for model diagnostics
b <- tidybayes::spread_draws(mod, b_Trt1)
# tibble:length = n_chains * n_draws (post warm-up)
# the mean(a$b_Trt1) is the same as the beta coefficient in summary(mod)
# thus, this is only the beta-coefficients' draws, not added to intercept
c <- tidybayes::add_linpred_draws(object = mod,
newdata = dat)
# tibble with prediction for each observation *n_chain*n_sample =>
# predicted posterior distribution on the link-level for each observation
# gives the same as model |> marginaleffects::predictions () |> get_draws() below
# if this goes through the inverse link function, we get at the E(x)
# connections:
fixef(mod) # reports the mean estimates for intercept and predictor Estimate Est.Error Q2.5 Q97.5
Intercept 2.14932242 0.03225532 2.0863278 2.2124241
Trt1 -0.07577729 0.04572413 -0.1659439 0.0119442mean(a$b_Trt1) # contains all samples of the predictor posterior[1] -0.07577729mean(b$b_Trt1) # the same as above[1] -0.07577729mean(c$.linpred) # generates predictions for each observation, i.e. intercept + effect of treatment, but still on the link-level[1] 2.109507mean(exp(c$.linpred)) = 8.2543903 is the average predicted number of seizures, now on the response level because the .linpred is back-converted through the inverse link function (in this case exp().
using marginaleffects package
The default of the marginaleffects package in terms of reporting is that for central tendencies, they take the median. This leads to sometimes slightly different numbers than tidybayes and brms. To change this, we set it to the mean, even thought I think the median is a sensible default.
options(marginaleffects_posterior_center = mean) # the default is the median, but later on, we compare with the mean
a <- marginaleffects::predictions(mod, type = "link")
# tibble with the estimate column giving the link-level estimates for the different groups. no variance, just intercept + b*x
# size: n_observations in model (i.e. all nas are out if you did not impute)
a <- marginaleffects::predictions(mod, type = "link", newdata = dat)
# tibble with a prediction for every observation, on the link level, no variance, just the intercept + b*x
# size: n_observations, with rows containing NA that were not part of the model
b <- get_draws(a) # tibble with n_chain*n_draw*n_observations,
# samples from the posterior distribution of the model
# depending on whether you specified newdata before, you will get predictions for all data, or all data that went into the model (without imputation, i.e without taking into account lines with missing data)
# this can be used to summarize the posterior distributionThe difference between the two estimates of the two conditions, unique(a$estimate) = 2.1493224, 2.0735451, is the same as the beta coefficient of the model itself:
The get_draws(a) however makes a bit more sense, its mean is exactly the same as the mean of brms::posterior_linpred(mod). The mean(exp(b$draw)) gives again the average predicted number of seizures, now on the response level because the draw on the link-level is back-converted through the inverse link function.
Then, to do summaries, there is the avg_predictions() and avg_comparisons() function. The comparisons()function takes a counterfactual approach by: 1) Modify the original dataset to fix for all observations, and compute predictions for every row. 2) Modify the original dataset to fix for all observations, and compute predictions for every row. 3) Calculate the average difference between the counterfactual predictions computed in steps 1 and 2.
See this website for more info.
## these two give the same. I am not sure why
# averaged over the predictions() from above:
marginaleffects::avg_predictions(mod, type = "link", variables = "Trt") # this gives the predictions for the two treatment conditions
Trt Estimate 2.5 % 97.5 %
0 2.15 2.09 2.21
1 2.07 2.01 2.14
Type: linkmarginaleffects::avg_comparisons(mod, type = "link", variables = "Trt") # this gives directly the difference between the treatment conditions
Estimate 2.5 % 97.5 %
-0.0758 -0.166 0.0119
Term: Trt
Type: link
Comparison: 1 - 0To test what they write on their webpage: ” type = “link”: Compute posterior draws of the linear predictor using the brms::posterior_linpred function.“, let’s compare the two directly:
a <- marginaleffects::predictions(mod, type = "link")
mean(a$estimate)[1] 2.109507mean(brms::posterior_linpred(mod))[1] 2.109507They are exactly the same.
Draws on the expected response level
In this part, we get draws that are back-transformed through the inverse link function. Given that in most brms models, per default, the linear part of the model predicts the (link-transformed) central tendency or expected value of the outcome distribution, the estimates that are back-transformed give only the expected values. Thus, the sigma term from brms, which could be the measurement error, is not taken into consideration. Therefore, credible intervals on this level are much narrower than in the next section on the predicted response level. As a consequence, we don’t get actual counts of seizures, but expected, or average counts, i.e. with decimal numbers.
using brms
Again, brms returns matrices, that are not super handy to work with, but these are the basis for the following functions. In this case, the _linpred(model, transform = TRUE) gives the same result, as the _epred(), but this is not always the case, see here, and here for why it will not always be the case (e.g. in lognormal models!)
a <- brms::posterior_linpred(mod, transform = TRUE)
# expected response, without sigma
# # matrix with n_samples x n_observations
b <- brms::posterior_epred(mod)
unique(a == b) [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10] [,11] [,12] [,13] [,14]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,15] [,16] [,17] [,18] [,19] [,20] [,21] [,22] [,23] [,24] [,25] [,26]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,27] [,28] [,29] [,30] [,31] [,32] [,33] [,34] [,35] [,36] [,37] [,38]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,39] [,40] [,41] [,42] [,43] [,44] [,45] [,46] [,47] [,48] [,49] [,50]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,51] [,52] [,53] [,54] [,55] [,56] [,57] [,58] [,59] [,60] [,61] [,62]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,63] [,64] [,65] [,66] [,67] [,68] [,69] [,70] [,71] [,72] [,73] [,74]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,75] [,76] [,77] [,78] [,79] [,80] [,81] [,82] [,83] [,84] [,85] [,86]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,87] [,88] [,89] [,90] [,91] [,92] [,93] [,94] [,95] [,96] [,97] [,98]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,99] [,100] [,101] [,102] [,103] [,104] [,105] [,106] [,107] [,108]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,109] [,110] [,111] [,112] [,113] [,114] [,115] [,116] [,117] [,118]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,119] [,120] [,121] [,122] [,123] [,124] [,125] [,126] [,127] [,128]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,129] [,130] [,131] [,132] [,133] [,134] [,135] [,136] [,137] [,138]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,139] [,140] [,141] [,142] [,143] [,144] [,145] [,146] [,147] [,148]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,149] [,150] [,151] [,152] [,153] [,154] [,155] [,156] [,157] [,158]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,159] [,160] [,161] [,162] [,163] [,164] [,165] [,166] [,167] [,168]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,169] [,170] [,171] [,172] [,173] [,174] [,175] [,176] [,177] [,178]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,179] [,180] [,181] [,182] [,183] [,184] [,185] [,186] [,187] [,188]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,189] [,190] [,191] [,192] [,193] [,194] [,195] [,196] [,197] [,198]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,199] [,200] [,201] [,202] [,203] [,204] [,205] [,206] [,207] [,208]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,209] [,210] [,211] [,212] [,213] [,214] [,215] [,216] [,217] [,218]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,219] [,220] [,221] [,222] [,223] [,224] [,225] [,226] [,227] [,228]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUE
[,229] [,230] [,231] [,232] [,233] [,234] [,235] [,236]
[1,] TRUE TRUE TRUE TRUE TRUE TRUE TRUE TRUEusing tidybayes()
a <- tidybayes::add_linpred_draws(object = mod, newdata = dat, transform = TRUE)
# tibbles with n_observation_n_chains*n_draws lines
b <-tidybayes::add_epred_draws(object = mod, newdata = dat)
c <-tidybayes::epred_draws(object = mod, newdata = dat)
unique(a$.linpred == b$.epred)[1] TRUEunique(a$.linpred == c$.epred)[1] TRUEunique(b$.epred == c$.epred)[1] TRUEIn this case, the three lines give the same result. But see here, and here for why it will not always be the case (e.g. in lognormal models!)
using marginaleffects
I use type = “response”: Compute posterior draws of the expected value using the brms::posterior_epred function. to get the expected values.
Get the draws:
# get the estimates on a unit level, conditional on all other predictors being average
a <- marginaleffects::predictions(mod, type = "response")
# tibble with prediction for each row of the dataset that was used fitting the model (is not the same as observation in the dataset if you did not impute)
b <- marginaleffects::predictions(mod, type = "response", newdata = dat) # prediction for each row of the dataset, can be original data or synthetic, new data.
# tibble with n_observatons (including those that had NA and were not part of the model if you did not impute)
c <- get_draws(b) # or (a), gives a tibble with n_chain*n_sample*n_observations rowsCompare conditions, overall numbers
marginaleffects::avg_predictions(mod, type = "response", by = "Trt")
Trt Estimate 2.5 % 97.5 %
0 8.58 8.06 9.14
1 7.96 7.44 8.47
Type: response# tibble with the mean of the predictions for each row of the dataset that was used fitting the model (is not the same as observation in the dataset if you did not impute, i.e. observations with a missing will not get a prediction)
marginaleffects::avg_predictions(mod, type = "response", newdata = dat) # prediction for each row of the dataset
Estimate 2.5 % 97.5 %
8.25 7.9 8.62
Type: response# tibble with one line, mean of n_observatons (including those that had NA and were not part of the model if you did not impute)Directly compare the two conditions
#compare the estimates by taking the difference
marginaleffects::comparisons(mod, by = "Trt", type = "response")
Trt Estimate 2.5 % 97.5 %
0 -0.626 -1.38 0.0983
1 -0.626 -1.38 0.0983
Term: Trt
Type: response
Comparison: 1 - 0marginaleffects::avg_comparisons(mod, by = "Trt", type = "response")
Trt Estimate 2.5 % 97.5 %
0 -0.626 -1.38 0.0983
1 -0.626 -1.38 0.0983
Term: Trt
Type: response
Comparison: 1 - 0Draws on the predicted response level
Here, we produce predictions for either our existing data frame, or new data. The predictions are now on the true outcome level, i.e. not expected number of seizures, but actual predictions of actual integers for each row of the data frame.
using brms
a <- brms::posterior_predict(mod)
# prediction of individual responses, with sigma
# matrix with n_samples (rows) x n_observations (cols)using tidybayes
a <- tidybayes::add_predicted_draws(object = mod, newdata = dat)
b <- tidybayes::predicted_draws(object = mod, newdata = dat)
unique(a$.prediction == b$.prediction)[1] FALSE TRUEThese are not always the same since there is now “sampling error” included in the predictions. Also, if you rerun each line, there will be different .predictions made.
using marinaleffects
a <- marginaleffects::predictions(mod, type = "prediction")
a <- marginaleffects::predictions(mod, type = "prediction", newdata = dat)
#type = "prediction": Compute posterior draws of the posterior predictive distribution using the brms::posterior_predict function.
b <- get_draws(a) # tibble with n_chain*n_sample*n_observations
a <- marginaleffects::avg_predictions(mod, type = "prediction")
# averaged over predictions() from aboveDirect comparison of outputs
In this part, I directly compare the output of tidybayesand marginaleffects, with by-hand computations of the estimates, to better understand what does what. I do it first continuing with the Poisson regression from above.
Poisson Regression
Poisson regressions use the log() as a link function, to ensure that the linear part of the model does not end up giving a lambda that is negative (which poisson()-family cannot handle). So, the model goes like this:
\(outcome \sim Poisson(lambda)\)
\(log(lambda) = a + bx\)
For a and b, we need to specify priors in Baeysian analysis, for example:
\(a \sim Normal(0,2)\)
\(b \sim Normsl(0,1)\)
We can use the marginaleffects-Package for prior predictive checks, or we can also use pp_check(). This is not of interest here.
Link-level comparison
First, I compare the output on the link-level. This is easiest because the posterior draws should directly translate to the results of summary(mod). I take the mean of the difference between the two treatment conditions, and as uncertainty-intervals, I take the 95%-equally-tailed-intervals (eti) around that mean. Another option is the 95% highest-density-interval (hdi). The eti is the default in marginaleffects and in fixef(), and it can easily be computed using the quantile() function. This choice for central tendency and uncertainty makes the results directly comparable betweent the packages / approaches.
Let’s look first at the increase of seizures between the two treatment groups with the different methods
link_comparison <- tibble(origin = character(),
estimate = numeric(),
lowerCI = numeric(),
upperCI = numeric())
## fixef baseline ----
fixef_row <- tibble(origin = "fixef(mod)", # from fixef as baseline
estimate = fixef(mod)[2,1],
lowerCI = fixef(mod)[2,3],
upperCI = fixef(mod)[2,4])
## tidybayes ----
# group_by(.draw, Trt)
# compute mean per draw
# pivot wider
# subtract group estimates within draw
# summarise
# correct way to do it: summarizing last
tidybayes_row <- mod |>
add_linpred_draws(newdata = dat,
transform = F) |> # stay on link level
group_by(.draw, Trt) |> # for each draw and level
summarise(.linpred = mean(.linpred), .groups = "drop") |> # take the mean => 2*4000 rows
tidyr::pivot_wider(names_from = Trt,
values_from = .linpred) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "tidybayes_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# if we don't summarize last, we get the wrong results:
# The following summarizes first and then does the subtraction, this produces much closer CIs
# There are two ways to this mistake.
#one
tidybayes_diffs <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = F) |> # not necesary, just for clarity
group_by(Trt) |>
summarise(mean_linpred = mean(.linpred),
lower_CI = quantile(.linpred, probs = 0.025),
upper_CI = quantile(.linpred, probs = 0.975)) |> # group expected values
summarise(across(2:4, ~ .[2] - .[1])) # group difference
tidybayes_row_w1 <- tibble(origin = "tidybayes_wrong1", # from fixef as baseline
estimate = tidybayes_diffs$mean_linpred,
lowerCI = tidybayes_diffs$lower_CI,
upperCI = tidybayes_diffs$upper_CI)
# tw0
a <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = F) |>
filter(Trt == 1) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
b <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = F) |>
filter(Trt == 0) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
tidybayes_diffs2 <- tibble(mean_linpred = a$y - b$y,
lower_CI = a$ymin - b$ymin,
upper_CI = a$ymax - b$ymax)
tidybayes_row_w2 <- tibble(origin = "tidybayes_wrong2", # from fixef as baseline
estimate = tidybayes_diffs2$mean_linpred,
lowerCI = tidybayes_diffs2$lower_CI,
upperCI = tidybayes_diffs2$upper_CI)
### marginaleffects -----
# per default, mariginaleffects uses the median as a central tendency
# https://marginaleffects.com/man/r/predictions.html#bayesian-posterior-summaries
# to make it comparable to the other things, change this to the mean:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti") # default, just to be explicit
# again here, there is a right and a wrong way to do it
# correct:
me_row_avg_pred_correct <- marginaleffects::predictions(mod,
newdata = dat,
type = "link") |>
get_draws() |>
group_by(drawid, Trt) |>
summarise(draws = mean(draw), .groups = "drop") |>
pivot_wider(names_from = Trt,
values_from = draws) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "marginaleffects::predictions_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# wrong: summarizing before subtracting
margEf_avgPred <- marginaleffects::avg_predictions(mod,
type = "link", # set to "link" or "response
by = "Trt") |> # grouped expected values
summarise(across(2:4, ~ .[2] - .[1])) # group differeneceWarning: There was 1 warning in `summarise()`.
ℹ In argument: `across(2:4, ~.[2] - .[1])`.
Caused by warning in `Ops.factor()`:
! '-' not meaningful for factorsme_row_avg_pred_w <- tibble(origin = "marginaleffects::avg_predictions_wrong", # from fixef as baseline
estimate = margEf_avgPred$estimate,
lowerCI = margEf_avgPred$conf.low,
upperCI = margEf_avgPred$conf.high)
# do it with the comparisons function
margEf_avgComp <- marginaleffects::avg_comparisons(mod,
type = "link", # set to "link" or "response
by = "Trt")
me_row_avg_comp <- tibble(origin = "marginaleffects::avg_comparisons_mean", # from fixef as baseline
estimate = margEf_avgComp$estimate,
lowerCI = margEf_avgComp$conf.low,
upperCI = margEf_avgComp$conf.high) Now, put together the table.
# A tibble: 8 × 4
origin estimate lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 fixef(mod) -0.0758 -0.166 0.0119
2 tidybayes_correct -0.0758 -0.166 0.0119
3 tidybayes_wrong1 -0.0758 -0.0793 -0.0754
4 tidybayes_wrong2 -0.0758 -0.0793 -0.0754
5 marginaleffects::predictions_correct -0.0758 -0.166 0.0119
6 marginaleffects::avg_predictions_wrong -0.0758 -0.0793 NA
7 marginaleffects::avg_comparisons_mean -0.0758 -0.166 0.0119
8 marginaleffects::avg_comparisons_mean -0.0758 -0.166 0.0119Interpreting the table is difficult because it is on the link-level.
Let’s look at the plots to see what is exactly equal:
tidybayes_draws <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = F) |>
ggplot(aes(x = .linpred, fill = Trt)) +
stat_halfeye() +
ylab("tidybayes::linpred")
marginaleff_pred_draws <- marginaleffects::predictions(model = mod, type = "link") |>
get_draws() |>
ggplot(aes(x = draw, fill = Trt)) +
stat_halfeye()+
ylab("marginaleffects::predictions")
cowplot::plot_grid(tidybayes_draws, marginaleff_pred_draws, nrow = 2,
labels = "poisson estimates on the link level")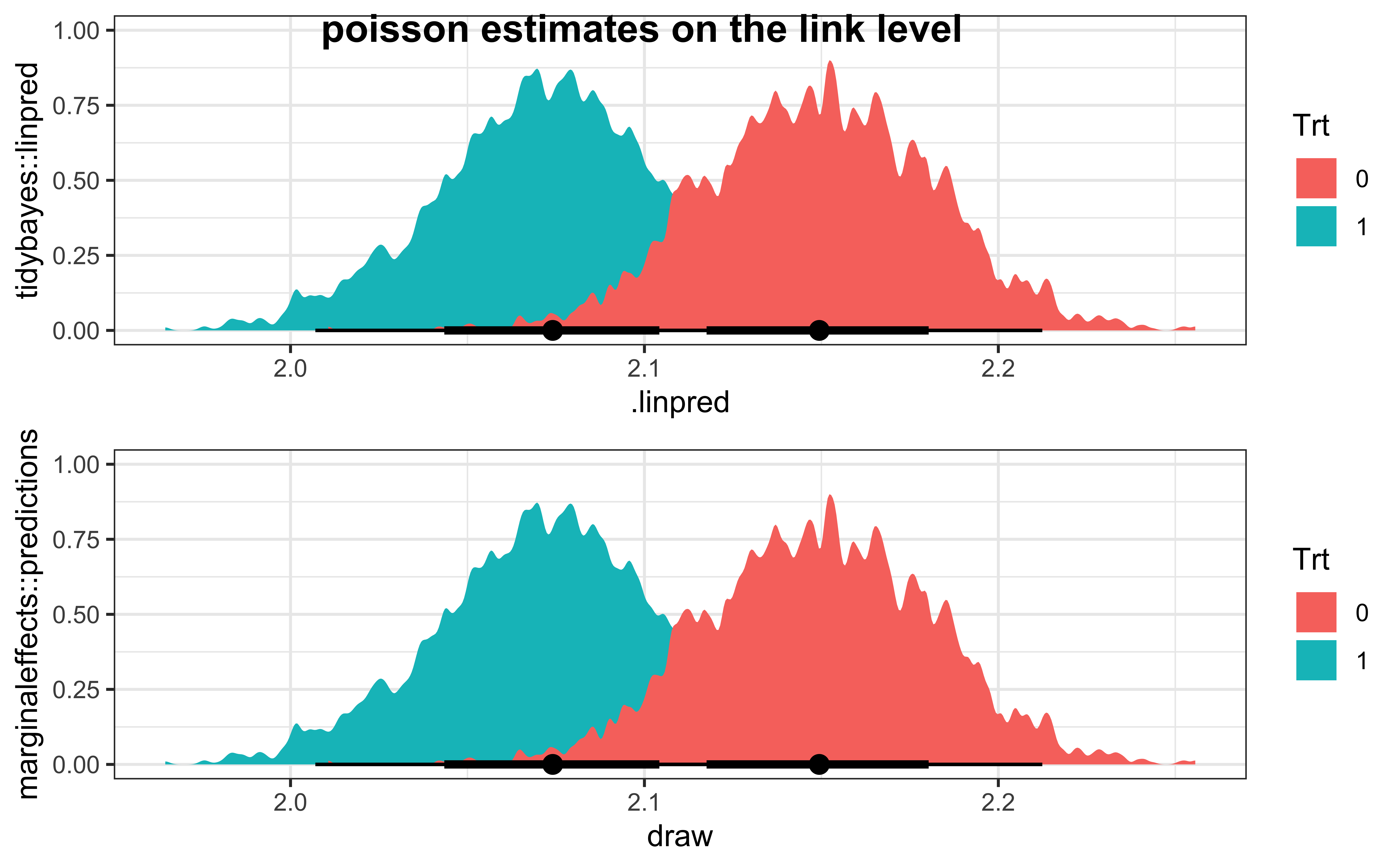
Expected response level comparison
Now, let’s look at the numbers on the response level. I compare between the back-transformed fixef() result and the results of tidybayes and marginaleffects. I am not entirely sure, if the back-transformation by hand is correct though.
response_comparison <- tibble(origin = character(),
estimate = numeric(),
lowerCI = numeric(),
upperCI = numeric())
## fixef baseline ----
# push link-level estimates through the inverse linkfunction
fixef_row <- tibble(origin = "fixef(mod)", # from fixef as baseline
estimate = exp(fixef(mod)[2,1] + fixef(mod)[1,1]) - exp(fixef(mod)[1,1]),
lowerCI = exp(fixef(mod)[2,3] + fixef(mod)[1,3]) - exp(fixef(mod)[1,3]),
upperCI = exp(fixef(mod)[2,4] + fixef(mod)[1,4]) - exp(fixef(mod)[1,4]))
## tidybayes ----
# group_by(.draw, Trt)
# compute mean per draw
# pivot wider
# subtract group estimates within draw
# summarise
# correct way to do it
tidybayes_row <- mod |>
add_linpred_draws(newdata = dat,
transform = T) |> # now on the response level
group_by(.draw, Trt) |> # for each draw and level
summarise(.linpred = mean(.linpred), .groups = "drop") |> # take the mean => 2*4000 rows
tidyr::pivot_wider(names_from = Trt,
values_from = .linpred) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "tidybayes_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# The following is wrong, summarizing first and then doing the comparison
# This summarizes first and then does the subtraction, this produces much closer CIs
# There are two ways to this mistake.
#one
tidybayes_diffs <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = T) |> # set T/F for response / link level
group_by(Trt) |>
summarise(mean_linpred = mean(.linpred),
lower_CI = quantile(.linpred, probs = 0.025),
upper_CI = quantile(.linpred, probs = 0.975)) |> # group expected values
summarise(across(2:4, ~ .[2] - .[1])) # group difference
tidybayes_row_w1 <- tibble(origin = "tidybayes_wrong1", # from fixef as baseline
estimate = tidybayes_diffs$mean_linpred,
lowerCI = tidybayes_diffs$lower_CI,
upperCI = tidybayes_diffs$upper_CI)
# twp
a <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = T) |>
filter(Trt == 1) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
b <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = T) |>
filter(Trt == 0) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
tidybayes_diffs2 <- tibble(mean_linpred = a$y - b$y,
lower_CI = a$ymin - b$ymin,
upper_CI = a$ymax - b$ymax)
tidybayes_row_w2 <- tibble(origin = "tidybayes_wrong2",
estimate = tidybayes_diffs2$mean_linpred,
lowerCI = tidybayes_diffs2$lower_CI,
upperCI = tidybayes_diffs2$upper_CI)
### marginaleffects -----
# per default, mariginaleffects uses the median as a central tendency
# https://marginaleffects.com/man/r/predictions.html#bayesian-posterior-summaries
# to make it comparable to the other things, change this to the mean:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti") # default, just to be explicit
# again here, there is a right and a wrong way to do it
# correct:
me_row_avg_pred_correct <- marginaleffects::predictions(mod,
newdata = dat,
type = "response") |>
get_draws() |>
group_by(drawid, Trt) |>
summarise(draws = mean(draw), .groups = "drop") |>
pivot_wider(names_from = Trt,
values_from = draws) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "marginaleffects::predictions_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# wrong: summarizing before subtracting
margEf_avgPred <- marginaleffects::avg_predictions(mod,
type = "response", # set to "link" or "response
variables = "Trt") |> # grouped expected values
summarise(across(2:4, ~ .[2] - .[1])) # group differenece
me_row_avg_pred_w <- tibble(origin = "marginaleffects::avg_predictions_wrong", # from fixef as baseline
estimate = margEf_avgPred$estimate,
lowerCI = margEf_avgPred$conf.low,
upperCI = margEf_avgPred$conf.high)
# do it with the comparisons function
margEf_avgComp <- marginaleffects::avg_comparisons(mod,
type = "response", # set to "link" or "response
variables = "Trt")
me_row_avg_comp <- tibble(origin = "marginaleffects::avg_comparisons_mean", # from fixef as baseline
estimate = margEf_avgComp$estimate,
lowerCI = margEf_avgComp$conf.low,
upperCI = margEf_avgComp$conf.high) # A tibble: 7 × 4
origin estimate lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 fixef(mod) -0.626 -1.23 0.110
2 tidybayes_correct -0.626 -1.38 0.0983
3 tidybayes_wrong1 -0.626 -0.614 -0.664
4 tidybayes_wrong2 -0.626 -0.614 -0.664
5 marginaleffects::predictions_correct -0.626 -1.38 0.0983
6 marginaleffects::avg_predictions_wrong -0.626 -0.614 -0.664
7 marginaleffects::avg_comparisons_mean -0.626 -1.38 0.0983The numbers in this table are mode interpretable. They are the expected difference in seizures between the treatment and the control group. However, the CI-numbers again are not exactly the same. the plots to see what is exactly equal:
tidybayes_draws <- tidybayes::add_linpred_draws(object = mod,
newdata = dat,
transform = T) |>
ggplot(aes(x = .linpred, fill = Trt)) +
stat_halfeye() +
ylab("tidybayes::linpred")
marginaleff_pred_draws <- marginaleffects::predictions(model = mod, type = "response") |>
get_draws() |>
ggplot(aes(x = draw, fill = Trt)) +
stat_halfeye()+
ylab("marginaleffects::predictions")
cowplot::plot_grid(tidybayes_draws, marginaleff_pred_draws, nrow = 2,
labels = "poisson estimates on the response level")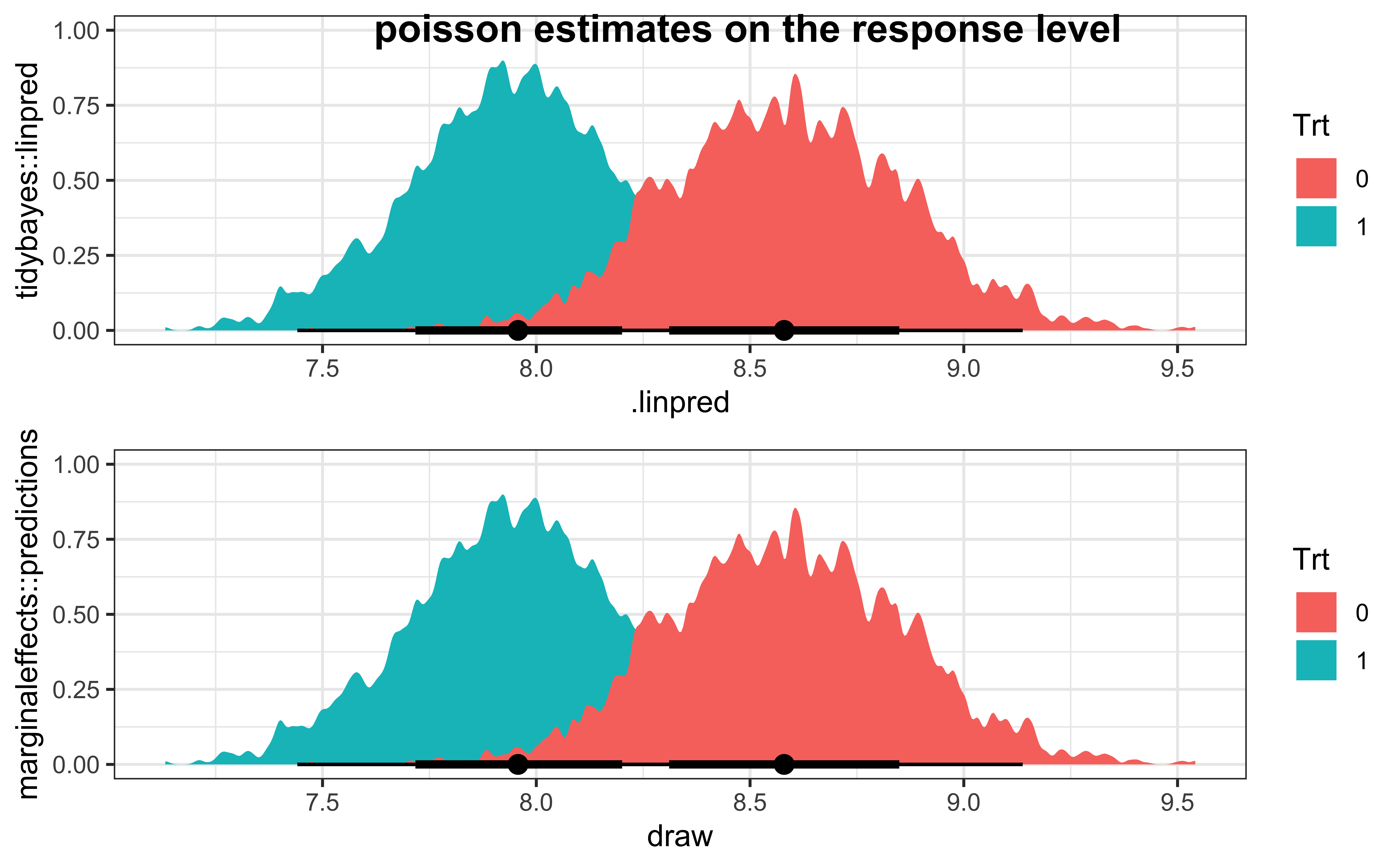
Bernoulli / Logistic regression
Now, let’s walk though another example, this time a logistic regression, with a new link function, the logit(). To make such a model, let’s dichotomize the count column into “many” and “not many” seizures, and predict it using the Trt variable again.
Link-level comparison
follow the same steps as above in the poisson model
link_comparison <- tibble(origin = character(),
estimate = numeric(),
lowerCI = numeric(),
upperCI = numeric())
## fixef baseline ----
fixef_row <- tibble(origin = "fixef(mod_logistic)", # from fixef as baseline
estimate = fixef(mod_logistic)[2,1],
lowerCI = fixef(mod_logistic)[2,3],
upperCI = fixef(mod_logistic)[2,4])
## tidybayes ----
# group_by(.draw, Trt)
# compute mean per draw
# pivot wider
# subtract group estimates within draw
# summarise
# correct way to do it
tidybayes_row <- mod_logistic |>
add_linpred_draws(newdata = dat_logistic,
transform = F) |> # stay on link level
group_by(.draw, Trt) |> # for each draw and level
summarise(.linpred = mean(.linpred), .groups = "drop") |> # take the mean => 2*4000 rows
tidyr::pivot_wider(names_from = Trt,
values_from = .linpred) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "tidybayes_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# This summarizes first and then does the subtraction, this produces much closer CIs
# There are two ways to this mistake.
#one
tidybayes_diffs <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = F) |> # set T/F for response / link level
group_by(Trt) |>
summarise(mean_linpred = mean(.linpred),
lower_CI = quantile(.linpred, probs = 0.025),
upper_CI = quantile(.linpred, probs = 0.975)) |> # group expected values
summarise(across(2:4, ~ .[2] - .[1])) # group difference
tidybayes_row_w1 <- tibble(origin = "tidybayes_wrong1", # from fixef as baseline
estimate = tidybayes_diffs$mean_linpred,
lowerCI = tidybayes_diffs$lower_CI,
upperCI = tidybayes_diffs$upper_CI)
# twp
a <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = F) |>
filter(Trt == 1) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
b <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = F) |>
filter(Trt == 0) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
tidybayes_diffs2 <- tibble(mean_linpred = a$y - b$y,
lower_CI = a$ymin - b$ymin,
upper_CI = a$ymax - b$ymax)
tidybayes_row_w2 <- tibble(origin = "tidybayes_wrong2", # from fixef as baseline
estimate = tidybayes_diffs2$mean_linpred,
lowerCI = tidybayes_diffs2$lower_CI,
upperCI = tidybayes_diffs2$upper_CI)
### marginaleffects -----
# per default, mariginaleffects uses the median as a central tendency
# https://marginaleffects.com/man/r/predictions.html#bayesian-posterior-summaries
# to make it comparable to the other things, change this to the mean:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti") # default, just to be explicit
# again here, there is a right and a wrong way to do it
# correct:
me_row_avg_pred_correct <- marginaleffects::predictions(mod_logistic,
newdata = dat_logistic,
type = "link") |>
get_draws() |>
group_by(drawid, Trt) |>
summarise(draws = mean(draw), .groups = "drop") |>
pivot_wider(names_from = Trt,
values_from = draws) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "marginaleffects::predictions_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# wrong: summarizing before subtracting
margEf_avgPred <- marginaleffects::avg_predictions(mod_logistic,
type = "link", # set to "link" or "response
variables = "Trt") |> # grouped expected values
summarise(across(2:4, ~ .[2] - .[1])) # group differenece
me_row_avg_pred_w <- tibble(origin = "marginaleffects::avg_predictions_wrong", # from fixef as baseline
estimate = margEf_avgPred$estimate,
lowerCI = margEf_avgPred$conf.low,
upperCI = margEf_avgPred$conf.high)
# do it with the comparisons function
margEf_avgComp <- marginaleffects::avg_comparisons(mod_logistic,
type = "link", # set to "link" or "response
variables = "Trt")
me_row_avg_comp <- tibble(origin = "marginaleffects::avg_comparisons_mean", # from fixef as baseline
estimate = margEf_avgComp$estimate,
lowerCI = margEf_avgComp$conf.low,
upperCI = margEf_avgComp$conf.high) Now, put together the table.
# A tibble: 7 × 4
origin estimate lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 fixef(mod_logistic) -0.482 -1.09 0.121
2 tidybayes_correct -0.482 -1.09 0.121
3 tidybayes_wrong1 -0.482 -0.544 -0.450
4 tidybayes_wrong2 -0.482 -0.544 -0.450
5 marginaleffects::predictions_correct -0.482 -1.09 0.121
6 marginaleffects::avg_predictions_wrong -0.482 -0.544 -0.450
7 marginaleffects::avg_comparisons_mean -0.482 -1.09 0.121And look at the plots to verify the output of tidybayes and marginaleffects are the same
tidybayes_draws <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = F) |>
ggplot(aes(x = .linpred, fill = Trt)) +
stat_halfeye() +
ylab("tidybayes::linpred")
marginaleff_pred_draws <- marginaleffects::predictions(model = mod_logistic, type = "link") |>
get_draws() |>
ggplot(aes(x = draw, fill = Trt)) +
stat_halfeye()+
ylab("marginaleffects::predictions")
cowplot::plot_grid(tidybayes_draws, marginaleff_pred_draws, nrow = 2,
labels = "logistic regression estimates on the link level")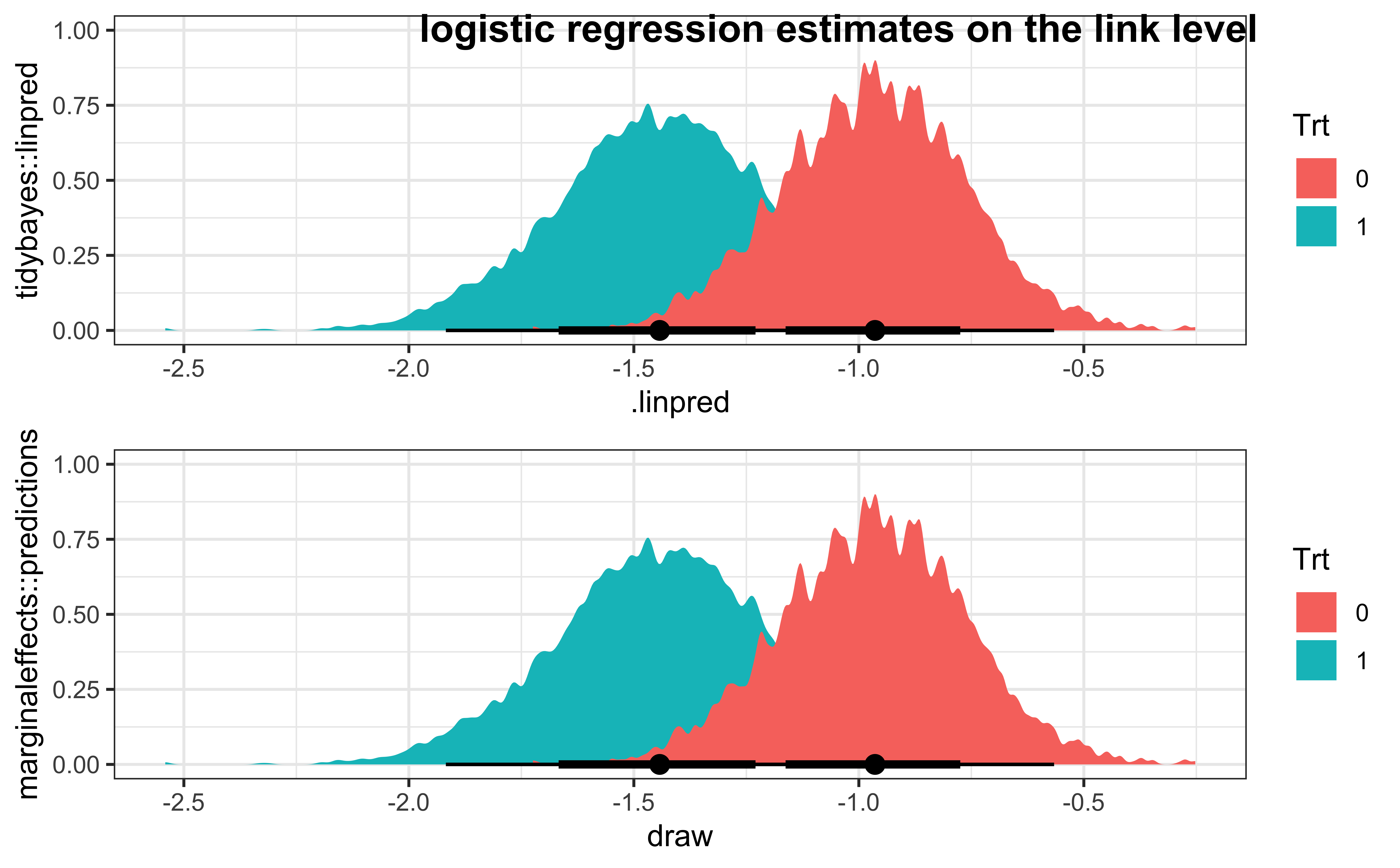
Expected response level comparison
Now, let’s look at the numbers on the response level. Because we will look at the estimated increase of the probability of having “many” seizures, the by-hand calculations become a bit complicated. Because of the non-linearity of the link-function, and the model-specification, the increase needs to be calculated by first adding Intercept and beta-coefficient, then inverse-linking it, then doing the same with only the intercept, then subtracting these two from one another. The numbers are a good fit for the conditional_effects() plot and therefore, I trust them.
response_comparison <- tibble(origin = character(),
estimate = numeric(),
lowerCI = numeric(),
upperCI = numeric())
## fixef baseline ----
# push link-level estimates through the inverse linkfunction
fixef_row <- tibble(origin = "fixef(mod)", # from fixef as baseline
estimate = boot::inv.logit(fixef(mod_logistic)[2,1] +
fixef(mod_logistic)[1,1]) -
boot::inv.logit(fixef(mod_logistic)[1,1]),
lowerCI = boot::inv.logit(fixef(mod_logistic)[2,3] +
fixef(mod_logistic)[1,3]) -
boot::inv.logit(fixef(mod_logistic)[1,3]),
upperCI = boot::inv.logit(fixef(mod_logistic)[2,4] +
fixef(mod_logistic)[1,4]) -
boot::inv.logit(fixef(mod_logistic)[1,4]))
## tidybayes ----
# group_by(.draw, Trt)
# compute mean per draw
# pivot wider
# subtract group estimates within draw
# summarise
# correct way to do it
tidybayes_row <- mod_logistic |>
add_linpred_draws(newdata = dat_logistic,
transform = T) |> # now on the response level
group_by(.draw, Trt) |> # for each draw and level
summarise(.linpred = mean(.linpred), .groups = "drop") |> # take the mean => 2*4000 rows
tidyr::pivot_wider(names_from = Trt,
values_from = .linpred) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "tidybayes_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# This summarizes first and then does the subtraction, this produces much closer CIs
# There are two ways to this mistake.
#one
tidybayes_diffs <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = T) |> # set T/F for response / link level
group_by(Trt) |>
summarise(mean_linpred = mean(.linpred),
lower_CI = quantile(.linpred, probs = 0.025),
upper_CI = quantile(.linpred, probs = 0.975)) |> # group expected values
summarise(across(2:4, ~ .[2] - .[1])) # group difference
tidybayes_row_w1 <- tibble(origin = "tidybayes_wrong1", # from fixef as baseline
estimate = tidybayes_diffs$mean_linpred,
lowerCI = tidybayes_diffs$lower_CI,
upperCI = tidybayes_diffs$upper_CI)
# twp
a <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = T) |>
filter(Trt == 1) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
b <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = T) |>
filter(Trt == 0) |>
pull(.linpred) |>
mean_qi(.linpred, .width = c(0.95))
tidybayes_diffs2 <- tibble(mean_linpred = a$y - b$y,
lower_CI = a$ymin - b$ymin,
upper_CI = a$ymax - b$ymax)
tidybayes_row_w2 <- tibble(origin = "tidybayes_wrong2", # from fixef as baseline
estimate = tidybayes_diffs2$mean_linpred,
lowerCI = tidybayes_diffs2$lower_CI,
upperCI = tidybayes_diffs2$upper_CI)
### marginaleffects -----
# per default, mariginaleffects uses the median as a central tendency
# https://marginaleffects.com/man/r/predictions.html#bayesian-posterior-summaries
# to make it comparable to the other things, change this to the mean:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti") # default, just to be explicit
# again here, there is a right and a wrong way to do it
# correct:
me_row_avg_pred_correct <- marginaleffects::predictions(mod_logistic,
newdata = dat_logistic,
type = "response") |>
get_draws() |>
group_by(drawid, Trt) |>
summarise(draws = mean(draw), .groups = "drop") |>
pivot_wider(names_from = Trt,
values_from = draws) |># prepare to take the difference
mutate(diff = `1` - `0`) |>
summarise(origin = "marginaleffects::predictions_correct",# now summarize over the draws
estimate = mean(diff),
lowerCI = quantile(diff, 0.025),
upperCI = quantile(diff, 0.975))
# wrong: summarizing before subtracting
margEf_avgPred <- marginaleffects::avg_predictions(mod_logistic,
type = "response", # set to "link" or "response
variables = "Trt") |> # grouped expected values
summarise(across(2:4, ~ .[2] - .[1])) # group differenece
me_row_avg_pred_w <- tibble(origin = "marginaleffects::avg_predictions_wrong", # from fixef as baseline
estimate = margEf_avgPred$estimate,
lowerCI = margEf_avgPred$conf.low,
upperCI = margEf_avgPred$conf.high)
# do it with the comparisons function
margEf_avgComp <- marginaleffects::avg_comparisons(mod_logistic,
type = "response", # set to "link" or "response
variables = "Trt")
me_row_avg_comp <- tibble(origin = "marginaleffects::avg_comparisons_mean", # from fixef as baseline
estimate = margEf_avgComp$estimate,
lowerCI = margEf_avgComp$conf.low,
upperCI = margEf_avgComp$conf.high) # A tibble: 7 × 4
origin estimate lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 fixef(mod) -0.0852 -0.124 0.0284
2 tidybayes_correct -0.0846 -0.192 0.0217
3 tidybayes_wrong1 -0.0846 -0.0740 -0.0963
4 tidybayes_wrong2 -0.0846 -0.0740 -0.0963
5 marginaleffects::predictions_correct -0.0846 -0.192 0.0217
6 marginaleffects::avg_predictions_wrong -0.0846 -0.0740 -0.0963
7 marginaleffects::avg_comparisons_mean -0.0846 -0.192 0.0217The numbers in this table indicate the increase in probability of having “many” seizures if one is in the treatment group compared to the control group. I have no clue as to why they are not the same.
And look at the plot again to verify that tidybayes and marginaleffects give the same
tidybayes_draws <- tidybayes::add_linpred_draws(object = mod_logistic,
newdata = dat_logistic,
transform = T) |>
ggplot(aes(x = .linpred, fill = Trt)) +
stat_halfeye() +
ylab("tidybayes::linpred")
marginaleff_pred_draws <- marginaleffects::predictions(model = mod_logistic, type = "response") |>
get_draws() |>
ggplot(aes(x = draw, fill = Trt)) +
stat_halfeye()+
ylab("marginaleffects::predictions")
cowplot::plot_grid(tidybayes_draws, marginaleff_pred_draws, nrow = 2,
labels = "logistic regression estimates on the response level")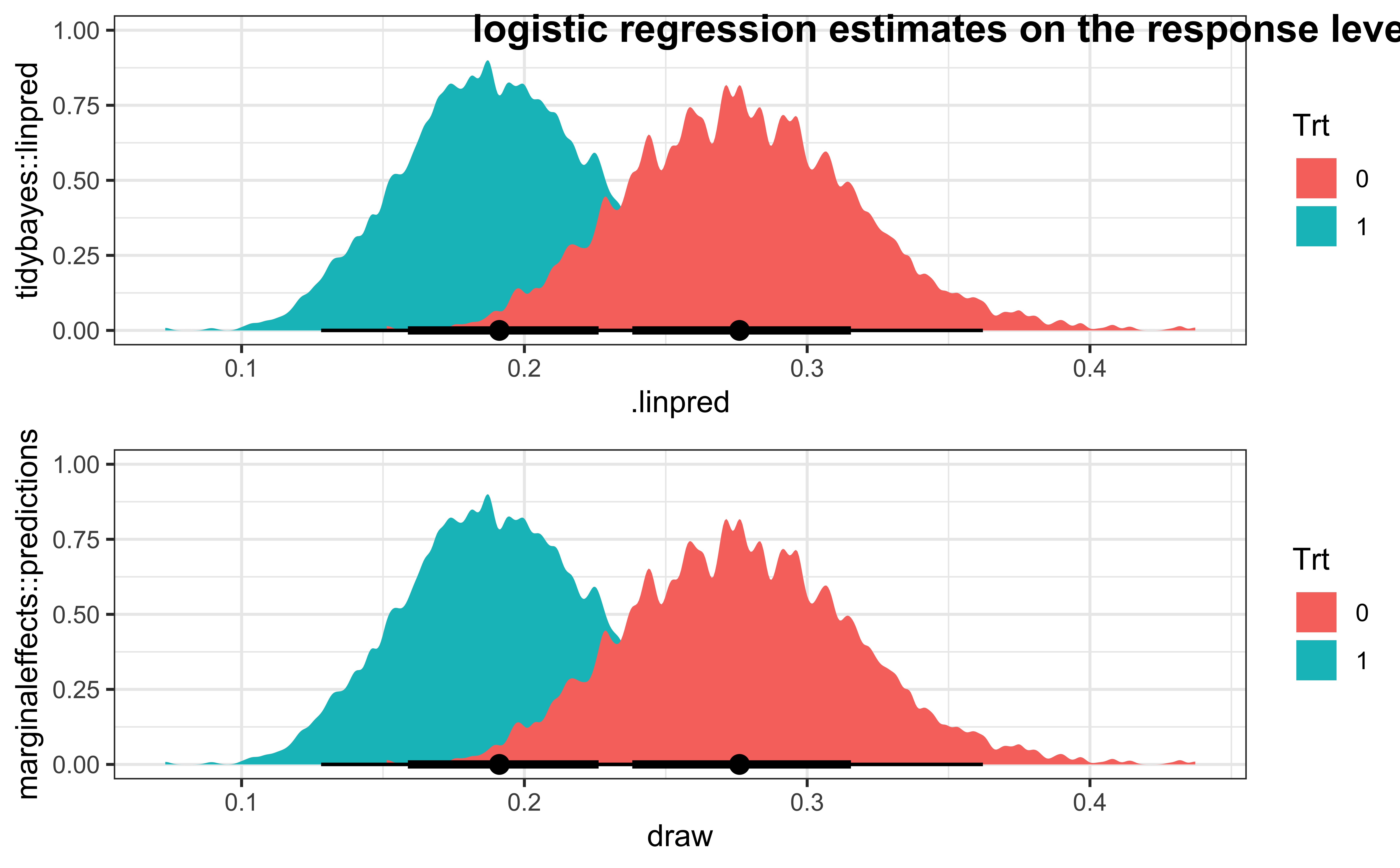
More realistic workflow - 2x2 interaction and continuous predictor
Lets say, we want to estimate the number of seizures by the interaction of Base and Treatment, controlled for age, i.e. the following model formula:
$count Base * Trt + zAge $
To simplify, I will dichotomize the Base variable with a median-split. I will show how to get at the estimated effects of the main effects, interactions, and how to make sense of the age predictor, which is continuous. I will get the draws for expected values, not the predicted ones.
realistic_data <- dat |>
mutate(Base_medianspl = case_when(Base >= 22 ~ "many",
Base < 22 ~ "few",
TRUE ~ "error"))
mod_realistic <- brm(data = realistic_data,
formula = bf(count ~ Base_medianspl * Trt + zAge),
family = poisson(),
file = "realistic_model")
draws <- mod_realistic |>
add_epred_draws(newdata = realistic_data) # this helps to let the draws not get too big, else they will take up too much RAM when you have several models in a project.Main effect treatment
First look at the average effect of the treatment, averaged over all Base values, and for a zAge of 0, i.e. average age.
`summarise()` has grouped output by '.draw'. You can override using the
`.groups` argument.# A tibble: 2 × 4
Trt est lowerCI upperCI
<fct> <dbl> <dbl> <dbl>
1 0 8.58 8.04 9.12
2 1 7.97 7.49 8.48### compute the difference between the two conditions
draws |>
group_by(.draw, Trt) |>
summarise(.epred = mean(.epred, na.rm = T)) |>
pivot_wider(names_from = Trt, values_from = .epred) |>
mutate(diff_Trt = `0` - `1`, .keep = "unused") |>
ungroup() |>
summarise(est = mean(diff_Trt),
lCI = quantile(diff_Trt, probs = 0.025),
uCI = quantile(diff_Trt, probs = 0.975))`summarise()` has grouped output by '.draw'. You can override using the
`.groups` argument.# A tibble: 1 × 3
est lCI uCI
<dbl> <dbl> <dbl>
1 0.606 -0.0866 1.33# have a look at the differences
draws |>
ggplot(aes(x = .epred, color = Trt)) +
geom_density()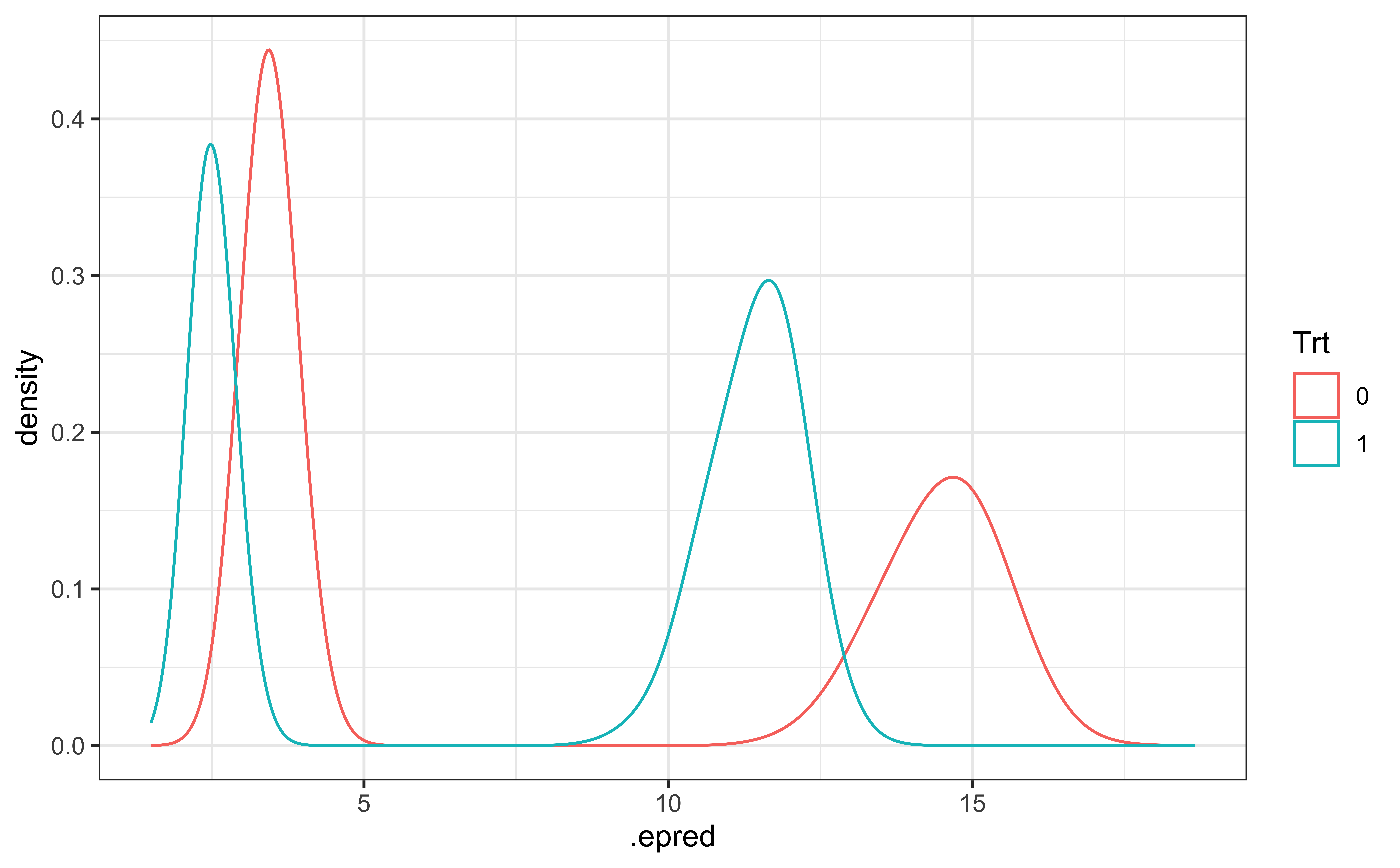
Interpretation: Patients with no treatment had an expected average seizure number of 8.59 (CI = [8.09; 9.13]), patients with treatment had an expected average seizure number of 7.97 (CI = [7.50; 8.46]). The difference between the two groups was 0.623 (CI = [-0.0180 1.38]).
This can also be calculated with the marginaleffects package:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti")
marginaleffects::predictions(mod_realistic,
newdata = realistic_data,
type = "response") |>
get_draws() |>
group_by(drawid, Trt) |>
summarise(estimate = mean(draw, na.rm = T)) |>
group_by(Trt) |>
summarize(est = mean(estimate),
lowerCI = quantile(estimate, probs = 0.025),
upperCI = quantile(estimate, probs = 0.975))`summarise()` has grouped output by 'drawid'. You can override using the
`.groups` argument.# A tibble: 2 × 4
Trt est lowerCI upperCI
<fct> <dbl> <dbl> <dbl>
1 0 8.58 8.04 9.12
2 1 7.97 7.49 8.48marginaleffects::avg_predictions(mod_realistic,
type = "link",
by = "Trt")
Trt Estimate 2.5 % 97.5 %
0 1.90 1.82 1.98
1 1.84 1.76 1.93
Type: linkMain effect Base
To get to the “effect” of the Baseline of seizures, averaged over both treatment groups, and for the zAge of 0.
`summarise()` has grouped output by '.draw'. You can override using the
`.groups` argument.# A tibble: 2 × 4
Base_medianspl est lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 few 3.03 2.71 3.37
2 many 12.7 12.1 13.3 ### compute the difference between the two groups
draws |>
group_by(.draw, Base_medianspl) |>
summarise(.epred = mean(.epred, na.rm = T)) |>
pivot_wider(names_from = Base_medianspl, values_from = .epred) |>
mutate(diff_base = many - few, .keep = "unused") |>
ungroup() |>
summarise(est = mean(diff_base),
lCI = quantile(diff_base, probs = 0.025),
uCI = quantile(diff_base, probs = 0.975))`summarise()` has grouped output by '.draw'. You can override using the
`.groups` argument.# A tibble: 1 × 3
est lCI uCI
<dbl> <dbl> <dbl>
1 9.65 8.97 10.4# have a look at the differences
draws |>
ggplot(aes(x = .epred, color = Base_medianspl)) +
geom_density()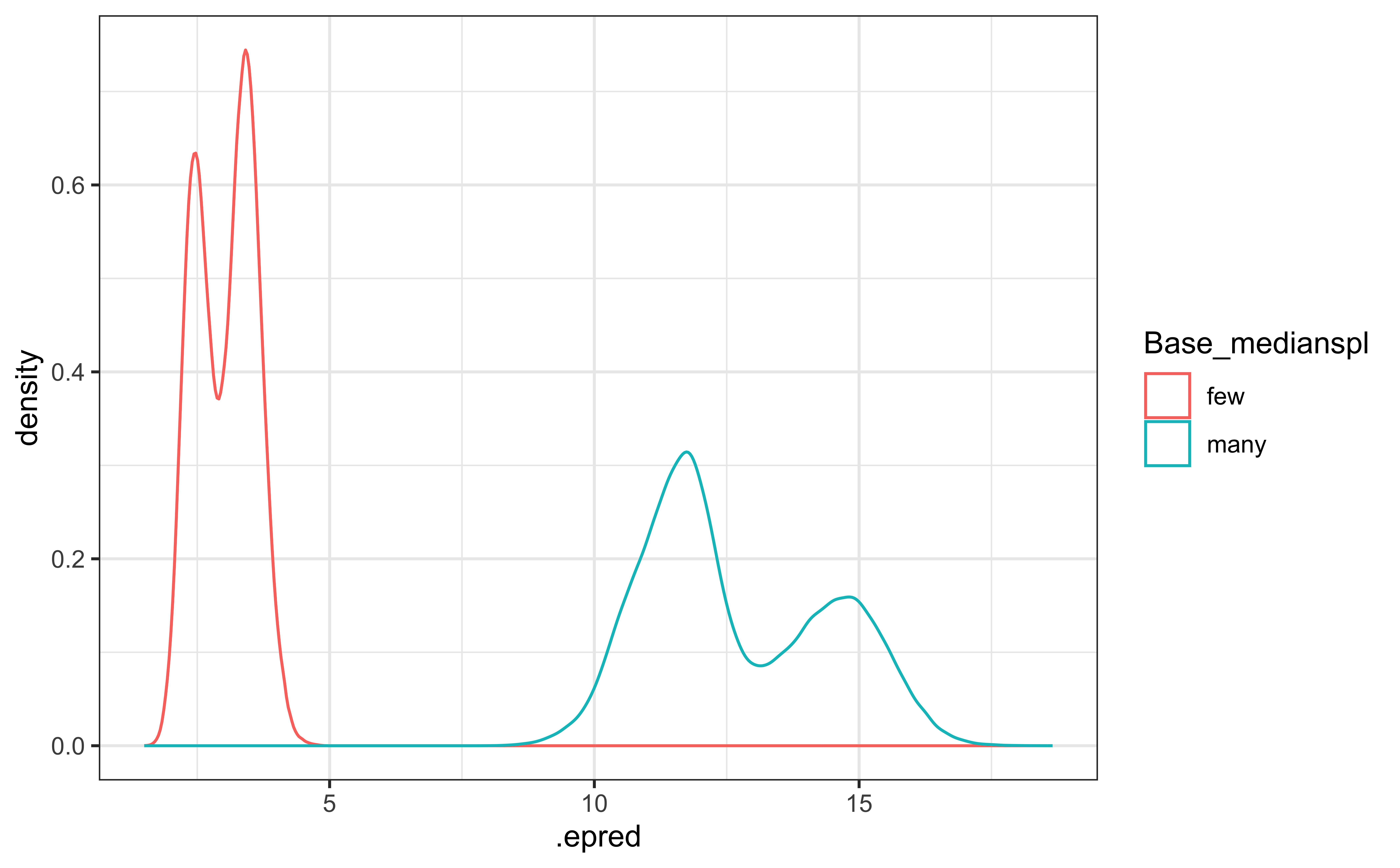
Patients starting with “few” seizures had an expected seizure number of 3.03 (CI = [2.71; 3.36]), those with “many” had an expected seizure number of 12.67 (CI = [12.06; 13.27]). The difference of expected seizures was 9.64 (CI = [8.95; 10.3]). Whether or not this difference is meaningful depends on the ROPE - the region of practical equivalence - that one should define beforehand. This is another topic in its own right.
This can also be calculated with the marginaleffects package:
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti")
marginaleffects::predictions(mod_realistic,
newdata = realistic_data,
type = "response") |>
get_draws() |>
group_by(drawid, Base_medianspl) |>
summarise(estimate = mean(draw, na.rm = T)) |>
group_by(Base_medianspl) |>
summarize(est = mean(estimate),
lowerCI = quantile(estimate, probs = 0.025),
upperCI = quantile(estimate, probs = 0.975))`summarise()` has grouped output by 'drawid'. You can override using the
`.groups` argument.# A tibble: 2 × 4
Base_medianspl est lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 few 3.03 2.71 3.37
2 many 12.7 12.1 13.3 marginaleffects::avg_predictions(mod_realistic,
type = "response",
by = "Base_medianspl")
Base_medianspl Estimate 2.5 % 97.5 %
few 3.03 2.71 3.37
many 12.68 12.07 13.29
Type: responseInteraction treatment and base
The interaction shows, whether the treatment-effect differs in the two clusters of patients, i.e. whether one patient group profits more from the treatment. See this site for more infos about the marginaleffectsside of things.
# difference of treatment effect in the two groups
draws |>
group_by(.draw, Base_medianspl, Trt) |>
summarise(.epred = mean(.epred, na.rm = T)) |>
group_by(Base_medianspl) |>
pivot_wider(names_from = Trt, values_from = .epred) |>
mutate(diff_Trt = `0` - `1`, .keep = "unused") |>
group_by(Base_medianspl) |>
summarize(est = mean(diff_Trt),
lowerCI = quantile(diff_Trt, probs = 0.025),
upperCI = quantile(diff_Trt, probs = 0.975))`summarise()` has grouped output by '.draw', 'Base_medianspl'. You can override
using the `.groups` argument.# A tibble: 2 × 4
Base_medianspl est lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 few 0.940 0.279 1.60
2 many 3.08 1.84 4.35### compute the difference of the treatment-differences between the two groups
draws |>
group_by(.draw, Base_medianspl, Trt) |>
summarise(.epred = mean(.epred, na.rm = T)) |>
group_by(Base_medianspl) |>
pivot_wider(names_from = Trt, values_from = .epred) |>
mutate(diff_Trt = `0` - `1`, .keep = "unused") |>
pivot_wider(names_from = Base_medianspl, values_from = diff_Trt) |>
mutate(diff_groups = many - few, .keep = "unused") |>
ungroup() |>
summarise(est = mean(diff_groups),
lCI = quantile(diff_groups, probs = 0.025),
uCI = quantile(diff_groups, probs = 0.975))`summarise()` has grouped output by '.draw', 'Base_medianspl'. You can override
using the `.groups` argument.# A tibble: 1 × 3
est lCI uCI
<dbl> <dbl> <dbl>
1 2.14 0.731 3.63# have a look at the differences
draws |>
ggplot(aes(x = .epred, color = Trt)) +
geom_density() +
facet_grid(.~Base_medianspl)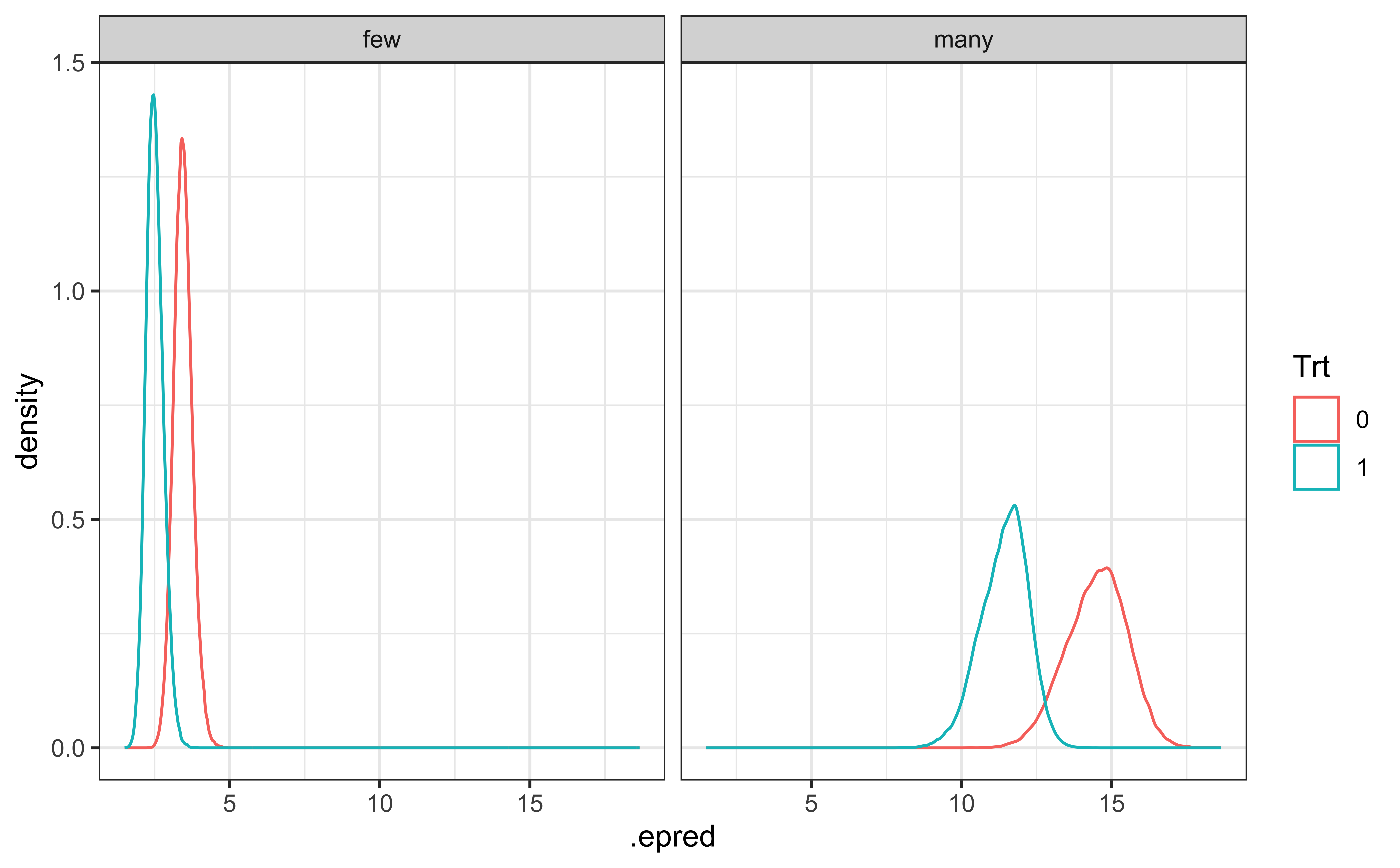
The treatment reduced expected seizure counts in both groups, but to different degrees. For people who started the treatment with “few” seizures, the reduction was 0.950 (CI = [0.295; 1.59]), for those with “many” seizures, it was 3.10 (CI = [1.83; 4.35]). The difference of the group-differences was 2.15 (CI = [0.725; 3.56]). Whether this difference is big enough to make us care should be determined with a ROPE. But this approach allows to formulate a ROPE on the outcome level and then compare the expected values to this ROPE.
This can also be calculated with the marginaleffects package, See here for more. For a contrast between predictions() and comparisons(), see here.
options("marginaleffects_posterior_center" = "mean", # not the default
"marginaleffects_posterior_interval" = "eti")
marginaleffects::predictions(mod_realistic,
newdata = realistic_data,
type = "response") |>
get_draws() |>
group_by(drawid, Base_medianspl, Trt) |>
summarise(.epred = mean(draw, na.rm = T)) |>
group_by(Base_medianspl) |>
pivot_wider(names_from = Trt, values_from = .epred) |>
mutate(diff_Trt = `0` - `1`, .keep = "unused") |>
group_by(Base_medianspl) |>
summarize(est = mean(diff_Trt),
lowerCI = quantile(diff_Trt, probs = 0.025),
upperCI = quantile(diff_Trt, probs = 0.975))`summarise()` has grouped output by 'drawid', 'Base_medianspl'. You can
override using the `.groups` argument.# A tibble: 2 × 4
Base_medianspl est lowerCI upperCI
<chr> <dbl> <dbl> <dbl>
1 few 0.940 0.279 1.60
2 many 3.08 1.84 4.35marginaleffects::avg_predictions(mod_realistic,
type = "response",
by = c("Base_medianspl", "Trt"))
Base_medianspl Trt Estimate 2.5 % 97.5 %
few 0 3.45 2.99 3.93
few 1 2.51 2.08 2.97
many 0 14.50 13.48 15.55
many 1 11.43 10.70 12.17
Type: responsemarginaleffects::avg_comparisons(mod_realistic,
variables = "Trt",
by = "Base_medianspl") # this gives a slightly different result, because it takes a counterfactual approach and contrasts all expected values if all rows had Treatment = 1 vs all rows if they had treatment = 0. But its in the same ballpark.
Base_medianspl Estimate 2.5 % 97.5 %
few -0.914 -1.57 -0.254
many -3.445 -4.79 -2.162
Term: Trt
Type: response
Comparison: 1 - 0marginaleffects::plot_predictions(mod_realistic,
type = "response",
by = c("Base_medianspl", "Trt")) # this give the same plot as: conditional_effects(mod_realistic)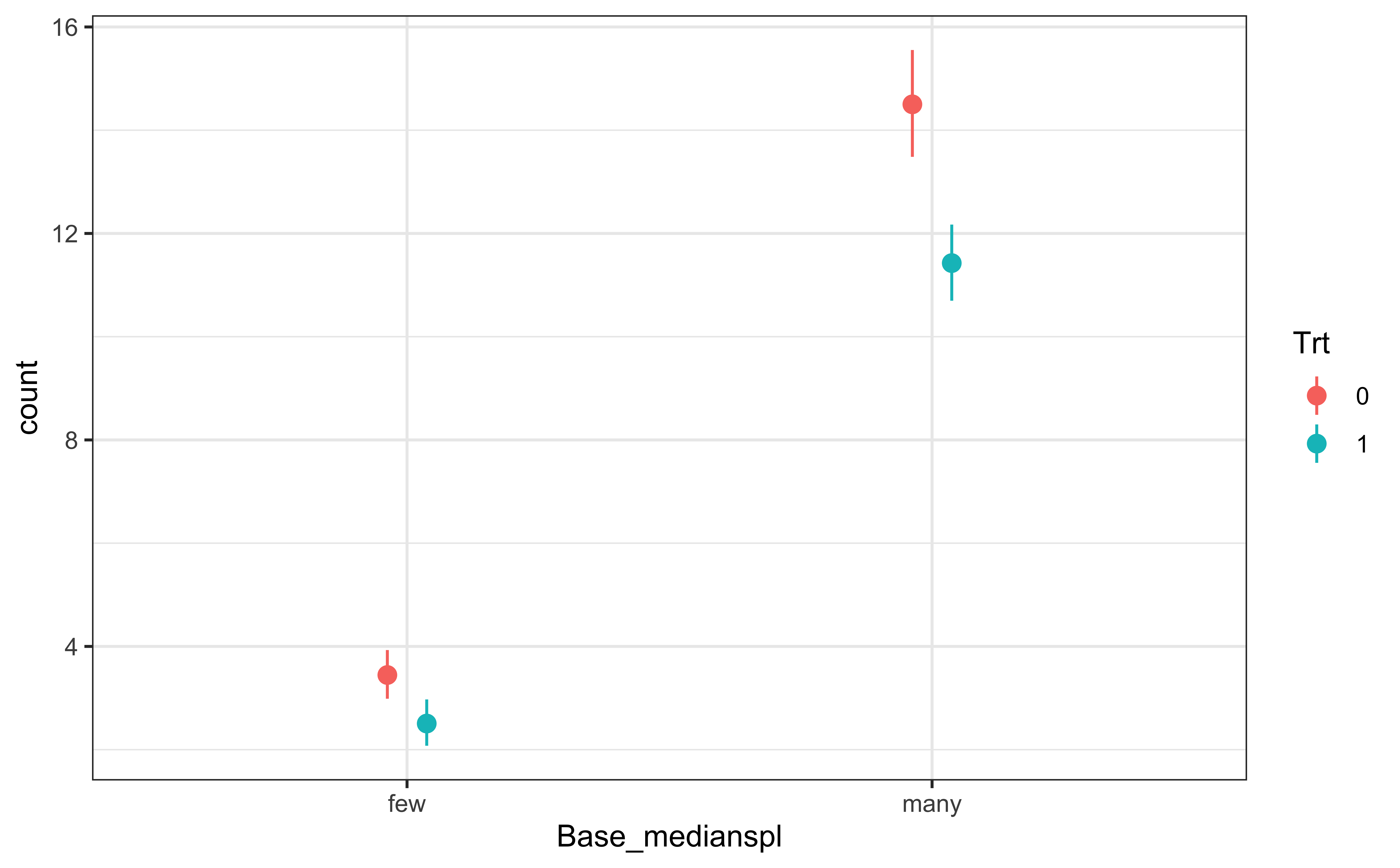
Slopes
The interpretation of the slope-parameter in a simple linear regression is simple: one unit change of the predictor translates to a slope-parameter-sized change in the expected value of dependent variable. In a generalized linear regression, this becomes more complicated: one unit change of the predictor translates to a slope-parameter-sized change in the link-function-transformed expected value of the dependent variable. This also means that a one unit increase on the link level does not lead to the same increase for all values of the predictor. Let’s illustrate this using the age-predictor. I cannot get my head around how to do this with tidybayes, so I will only do it with marginaleffects, which provides very nice functions. See here for more infos.
First, let’s look at the overall effect of age on seizures. We can see that seizures decrease in older patients. This is true for both Base-groups and for both treatment groups.
plot_predictions(mod_realistic,
variables = "zAge",
condition = c("zAge", "Trt", "Base_medianspl"), type = "response")+
labs(title = "How age impacts the seizures.",
subtitle = "For the different conditions.")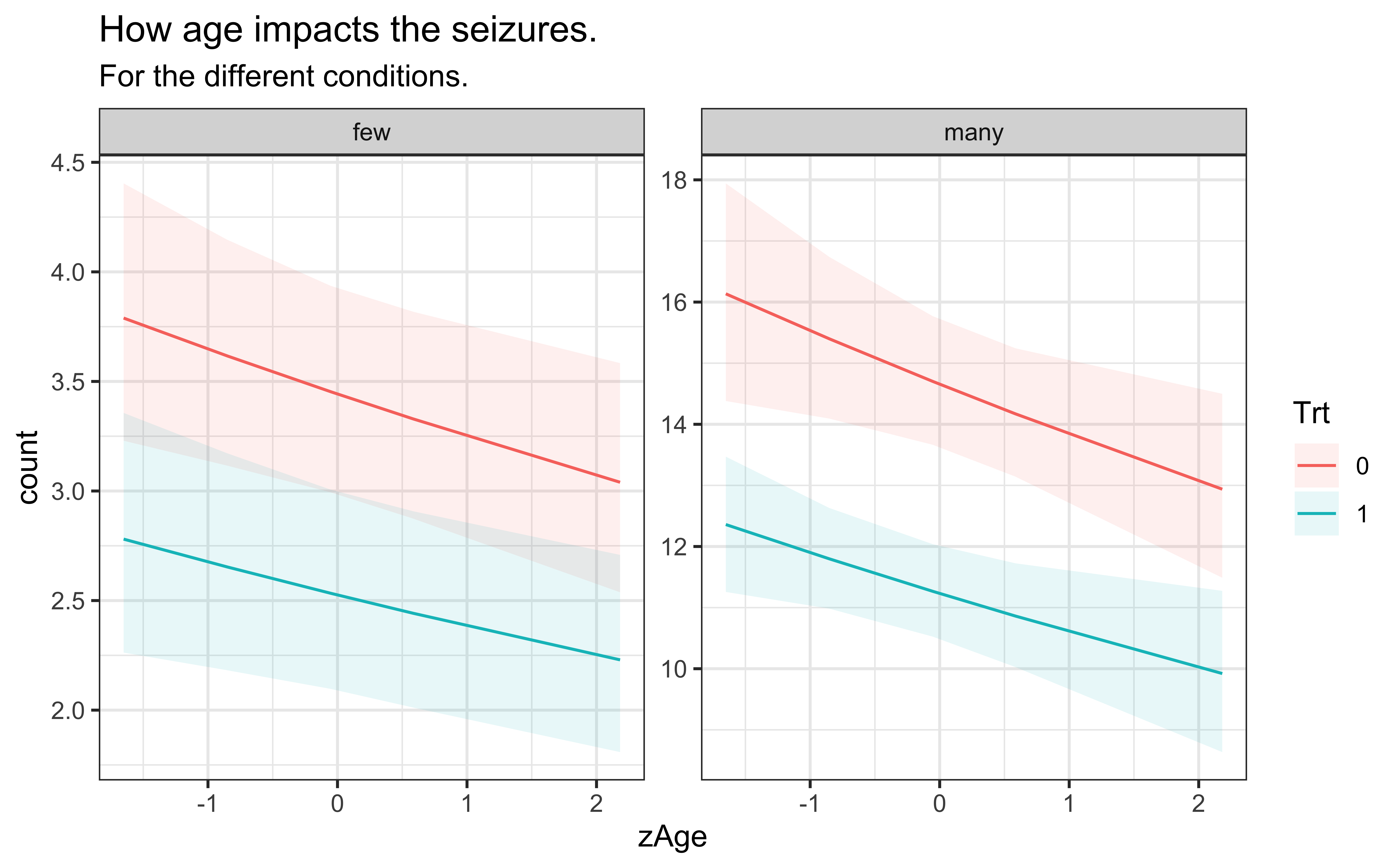
Next, lets look at how the slope (or the effect of age on seizures) changes over the different ages.
plot_slopes(mod_realistic,
variables = "zAge",
condition = c("zAge", "Trt", "Base_medianspl"), type = "response") +
labs(title = "How the effect of age on the seizures changes, as age increases.",
subtitle = "For the different conditions.")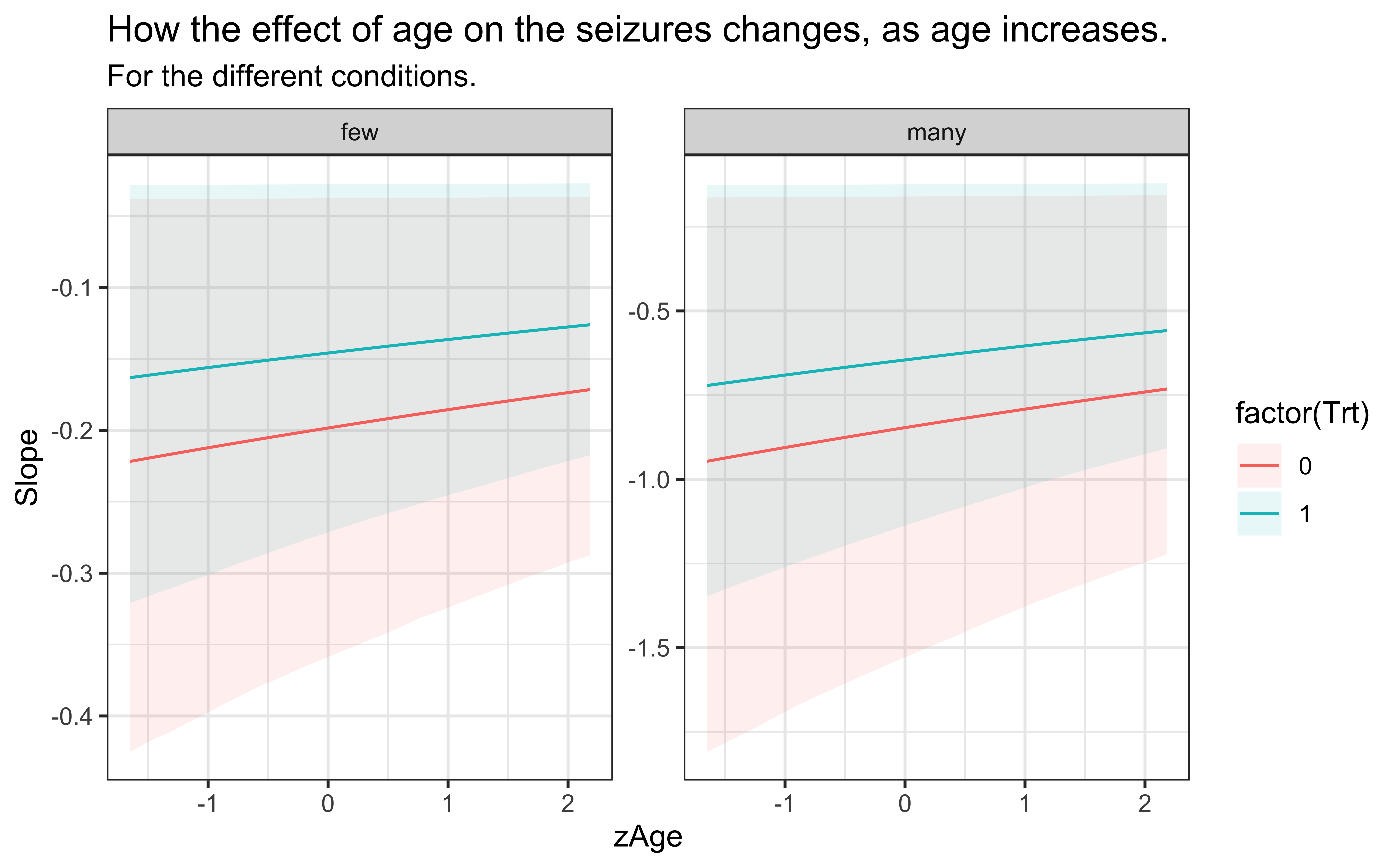
This plot becomes a line if we go to the link-level, which makes sense: the slope is the same for all ages on the link level, because it is a line.
plot_slopes(mod_realistic,
variables = "zAge",
condition = c("zAge", "Trt", "Base_medianspl"), type = "link") 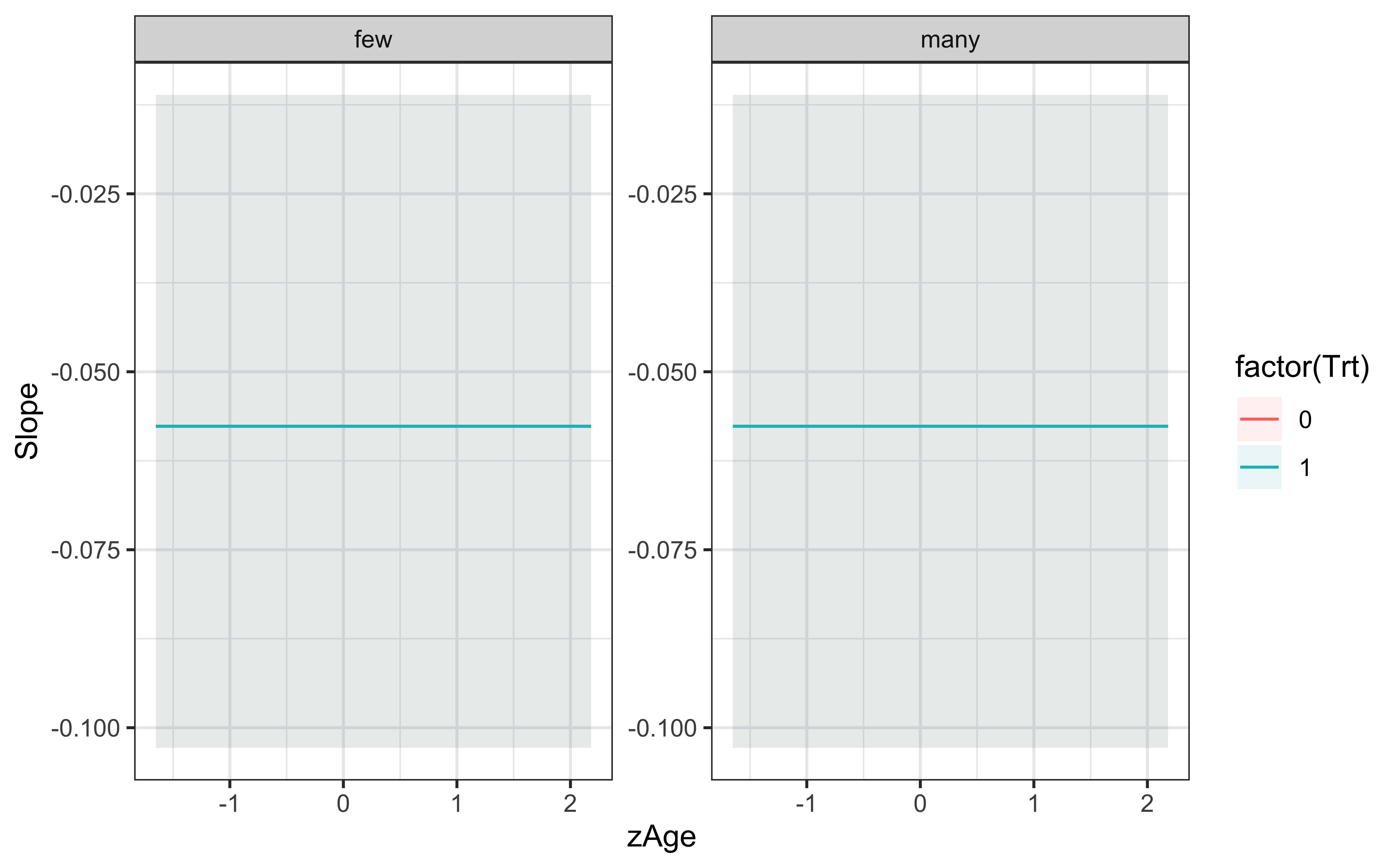
But to describe the results, we are interested in the average, or marginal effect of age on expected seizures. To get to this, we first plot:
mfx <- slopes(mod_realistic,
variables = "zAge",
newdata = datagrid(Trt = 0:1, Base_medianspl = c("many", "few")),
type = "response") |>
get_draws()
ggplot(mfx, aes(x = draw, fill = factor(Trt))) +
stat_halfeye(slab_alpha = .5) +
labs(x = "Marginal Effect of Age on Seizures",
y = "Posterior density",
fill = "Treatment") +
facet_grid(.~Base_medianspl)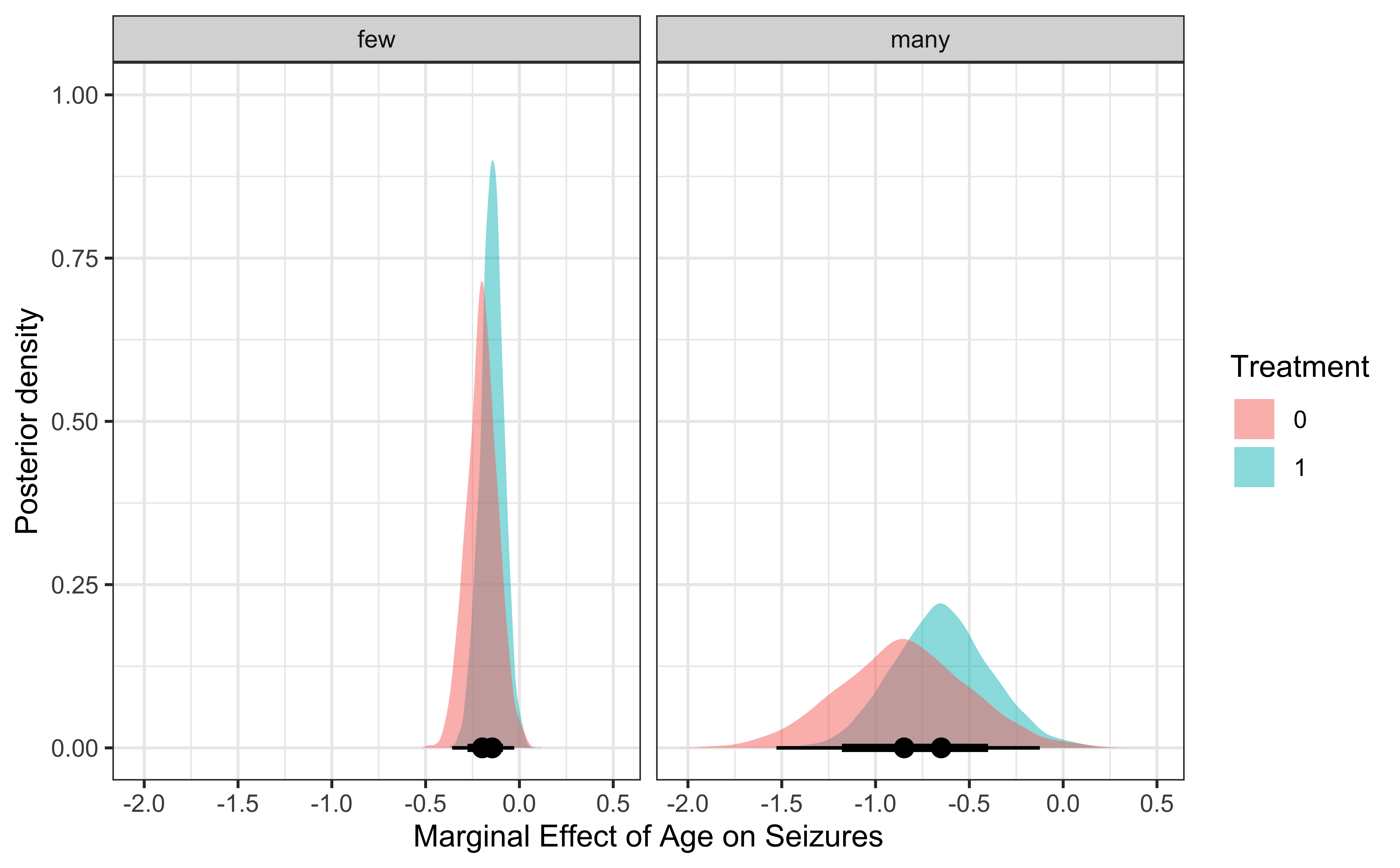
The best-off patients (in terms of age benefit, not in general) are the ones with many seizures to begin with and no treatment: Their age reduces their seizures the most per year of life (up to 1.5 seizures for example). To get the numbers of the average slope, we can do:
avg_slopes(mod_realistic, type = "response", variables = "zAge")
Estimate 2.5 % 97.5 %
-0.476 -0.853 -0.0902
Term: zAge
Type: response
Comparison: dY/dXAnd to get that in a plot:
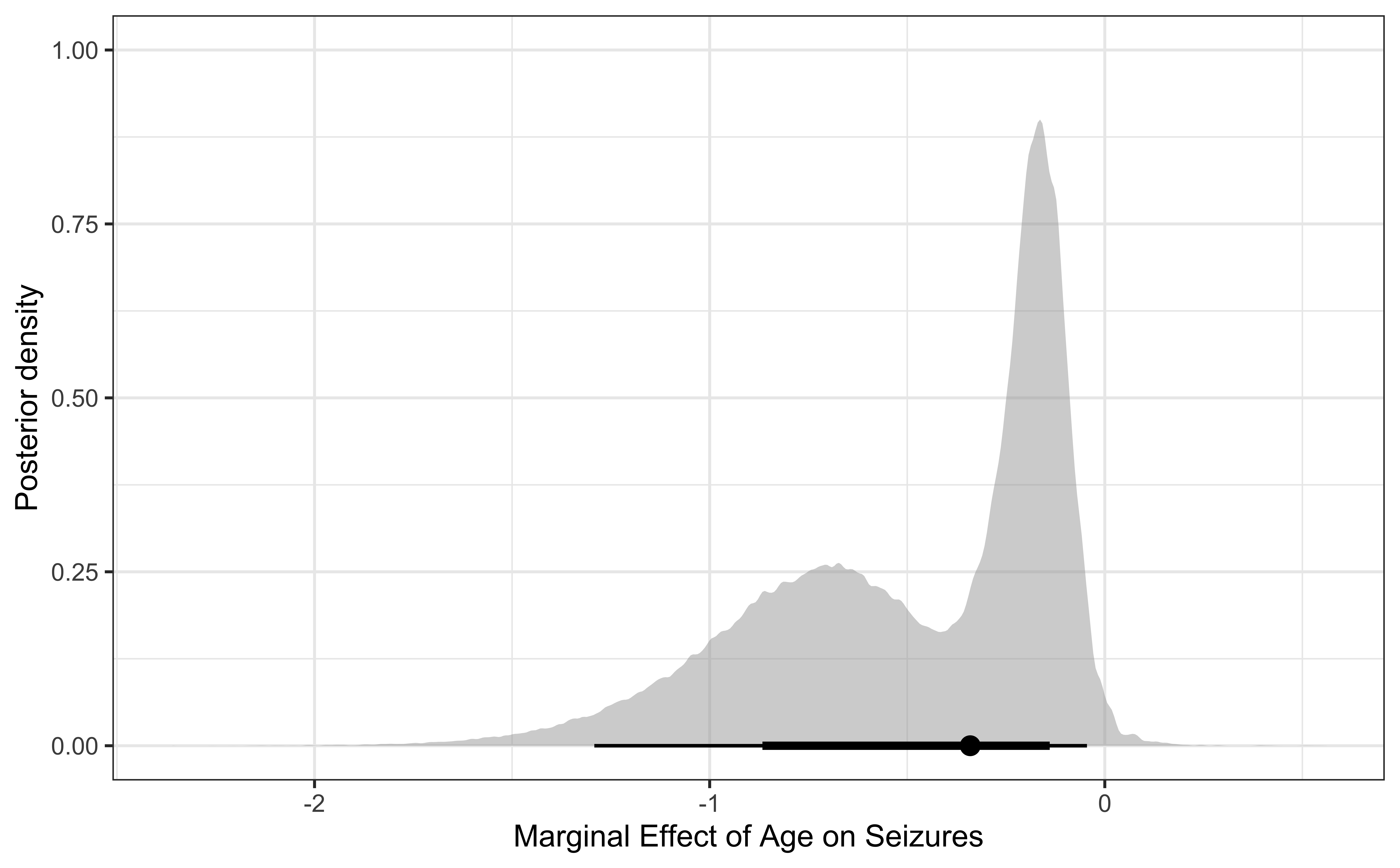
This plot makes it clear that the single slope for age is limited in its usefullness: for all combinations of treatment and baseline seizures, age decreases the number of seizures, but it makes a big difference for the age-effect whether a patient had many or few seizures to begin with:
ggplot(mfx, aes(x = draw, fill = Base_medianspl)) +
stat_halfeye(slab_alpha = .5) +
labs(x = "Marginal Effect of Age on Seizures",
y = "Posterior density",
fill = "Treatment") 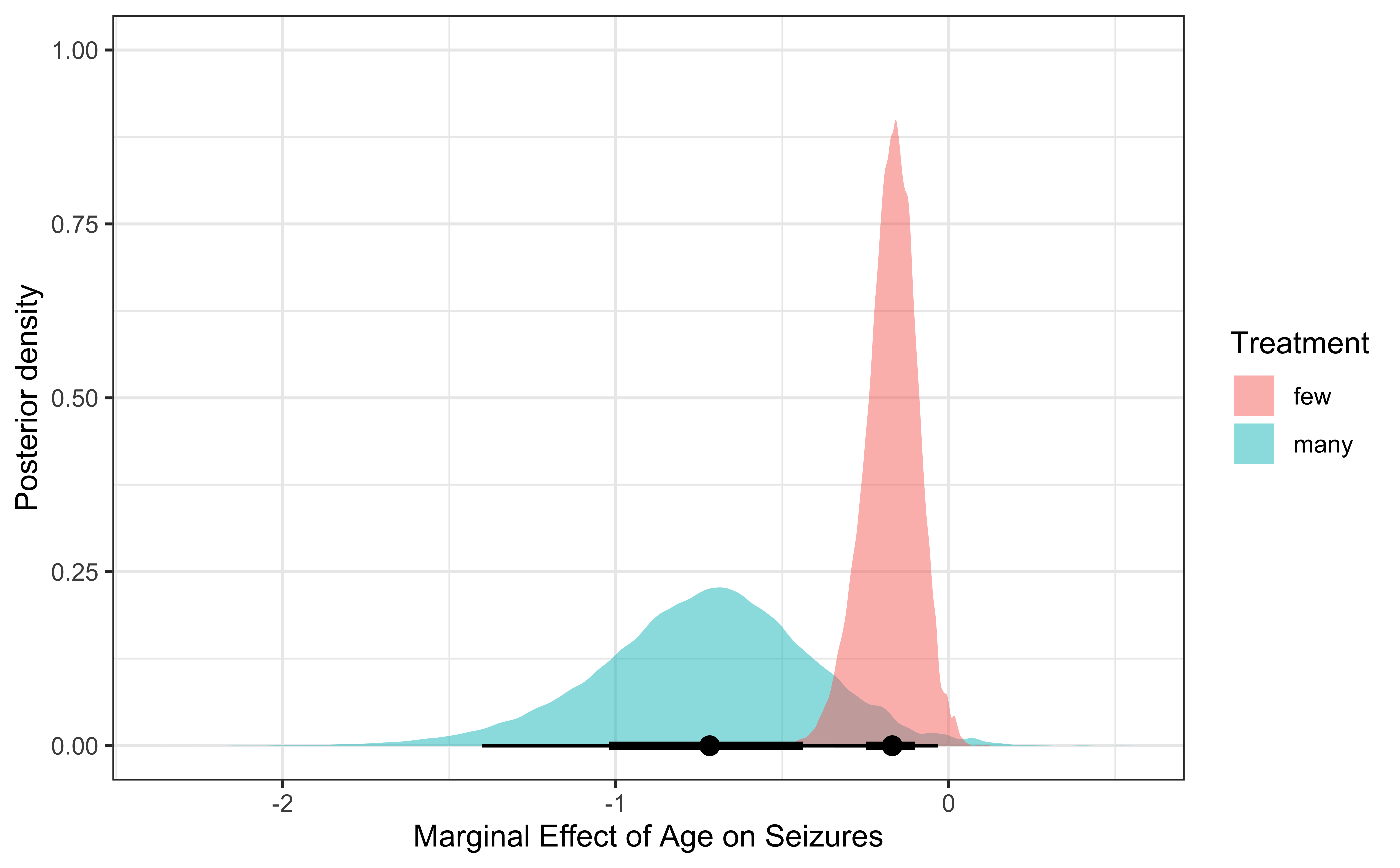
tl;dr
Both tidybayes and mariginaleffects have nice functions to get to the estimates of generalized linear models, both on the link, the expected and the predicted response levels. They can be made to give the same results. The most important thing is to always summarize last, work as long as possible with the draws.
However, the calculation of the credible intervals is not always consistent across link and response levels. The credible intervals of marginaleffects::avg_comparisons(), marginaleffects::predictions(), and tidybayes::linpred() are identical to the CIs of fixef(model) on the link-level. But the central tendencies and CIs are slightly off for the response-level estimates, which is because the link-function leads to non-linear behaviour. Essentially, because:
\(E(inverse.linkfunction(η)) \neq inverse.linkfunction(E(η))\)
if the link function is not the identity function.
The central tendency of marginaleffects is by default the median, which is why, on the link level, it gives a slightly different number than fixef() if this is not changed with: options("marginaleffects_posterior_center" = "mean", "marginaleffects_posterior_interval" = "eti"). The eti however is the default.
Caveat: I have only tried this with models with one predictor. These things might change as there are more predictors, because then, things get more complicated.
| Level | tidybayes | marginaleffects |
|---|---|---|
| Link-level | add_linpred_draws(model) |
predictions(model, type = "link") |
| Expected-response level | add_epred_draws(model) |
predictions(model, type = "response") |
| Predicted-response level | add_predicted_draws(model) |
predictions(model, type = "prediction") |
| Slopes | Don’t know yet how to do this |
avg_slopes(model, type = "link" / "response" / "prediction", variables = "continuous_variable_name")
|
| And then? | Then compare and summarize as needed |
avg_predictions(model,type = "link" / "response" / "prediction", by = c("var1", "var2")) or get_draws(), compare and summarize as needed. |
| Central Tendency & CI | get central tendency and CI by hand using quantile() for ETI, or hdi() for HDI. |
options("marginaleffects_posterior_center" = "median","marginaleffects_posterior_interval" = "eti")
|